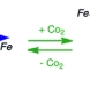New protein discovered gives insights to iron's fate underground
(Phys.org) -- It's almost an evil twin story: a protein that steals electrons from iron in one microbe looks a lot like one that adds electrons in another microbe, according to scientists at Pacific Northwest National Laboratory and the University of East Anglia. Their survey of the genes of common groundwater bacterium Sideroxydans lithotrophicus ES-1, which removes electrons from iron, revealed that it contained genes in common with Shewanella oneidensis MR-1, which adds electrons to iron.
Their results contribute to understanding of the molecular mechanisms by which microorganisms change the electron configuration of iron and, thus, change its mobility. The research was published in Frontiers in Microbiological Chemistry.
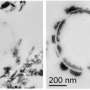
"Recent studies indicate that aerobic Fe(II)-oxidizing bacteria, FeOB, would play a key role in niches having low levels of oxygen concentration, where microbial Fe(II)-oxidation can compete with the chemical oxidation of Fe(II)," said PNNL biogeochemist Dr. Juan Liu, first author of the study paper.
Science has realized the importance of microorganisms in research on processes such as carbon sequestration, the generation of new energy sources, and the movement and ultimate resting place of contaminants. Scientists are interested in the oxidation state, or loss of electrons, of iron because it dramatically affects the metal's solubility in water, in which electron transfer proteins play critical roles. In contrast to Fe(II), trivalent iron, Fe(III), is not water soluble.
The difference in solubility between Fe(II) and Fe(III) also means that iron acquisition tends to be much more of a problem for organisms that use oxygen than for those that don't, because anaerobic environments favor the more soluble Fe(II).
"We have shown the generality of these reaction mechanisms in metal oxidizing and reducing bacteria," said Dr. Liang Shi, a PNNL microbiologist who led the study. "Whether it's Fe(II) or Fe(III), iron's solubility affects its accessibility to microorganisms. To access these different phases of Fe, some microorganisms seem to adopt a common mechanism."
The research team used integrated bioinformatics analyses, gene cloning, protein expression and purification, and in vitro and in vivo characterization on the ES-1 sample. Characterizations were conducted by using different experimental tools, such as complementation testing in MR-1, stopped-flow kinetics, and protein film cyclic voltammetry.
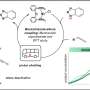
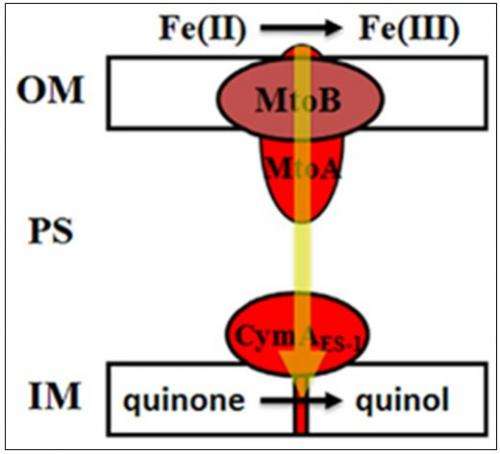
They found a three-gene cluster encoding homologs of MR-1. The MR-1 genes are called MtrA, MtrB, and CymAt, so to distinguish the ES-1 genes, the scientists named them MtoAB and CymAES-1.
From this cluster, they identified, purified, and characterized MtoA, a hemoprotein important for extracellular electron transfer reactions. MtoA is the first member of its specific family of proteins to be purified and characterized from an Fe(II)-oxidizing bacterium.
The scientists are planning to investigate to what extent these proteins are distributed among other Fe(II)-oxidizing bacteria.
More information: J Liu, et al.. 2012. "Identification and Characterization of MtoA: A Decaheme c-type Cytochrome of the Neutrophilic Fe(II)-oxidizing Bacterium Sideroxydans lithotrophicus ES-1." Frontiers in Microbiological Chemistry 3:37. DOI:10.3389/fmicb.2012.00037
Provided by Pacific Northwest National Laboratory