Enzyme's 'hop' offers tips on epigenetic gene expression
(Phys.org) -- UC Santa Barbara researchers' discovery of a variation of an enzyme's ability to "hop" as it moves along DNA, modifying the genetic material of a bacteria — and its physical capability and behavior — holds much promise for biomedical and other scientific applications. Their results are published in a paper titled "Proximal recognition sites facilitate intrasite hopping by DNA adenine methyltransferase: a mechanistic exploration of epigenetic gene regulation," in the Journal of Biological Chemistry.
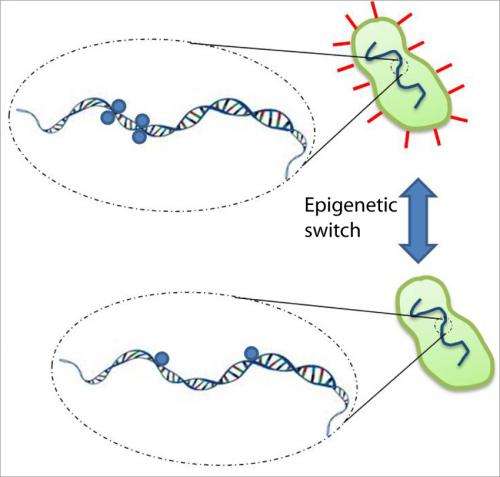
The E. coli bacteria's adaptive mechanism allows it to change its phenotype — its observable characteristics — according to its environment. For example, if it senses a need to find food, to stick to the tissues of its host organism, or to reproduce, the bacteria will form pili, or hairlike structures, on its surface, to allow it to move, stick or pass genetic material.
"We're trying to figure out what is it in the cell that's driving those changes," said Adam Pollak, first author of the paper.
The formation of these pili is driven by an epigenetic mechanism — a "tagging" done by the enzyme DNA adenine methyltransferase (Dam), which acts on a specific sequence of DNA, called GATC sites (Guanine-Adenine-Thymine-Cytosine). The tagging signals the formation of these appendages — a mechanism similar to that in humans, where tagging directs the formation of tissues for different organs from the same DNA. This tagging is part of a broader field, called epigenetics, where modifications made to the genome are heritable and regulate the expression of genes.
Where the prevailing belief used to be that the enzyme Dam slid down only one side of the bacteria's double-helixed DNA looking for these GATC sites, according to the researchers, Dam can actually "hop" to one or more such sites on both sides of the double helix.
"It moves along, finds a site, and methylates that; but it turns around, reorients itself, and methylates the other side," said Norbert Reich, UC Santa Barbara professor of chemistry and biochemistry.
Using several strands of genetically engineered DNA of various lengths and differing distances between the sites of methylation, the researchers found that the hopping of Dam may occur more often, depending on the clustering of sites; e.g., it is more likely to occur when two sites are within 10 to 200 base pairs of each other. Clustered GATC sites are strongly associated with gene regulation, while an isolated GATC site on the double helix is associated with the copying of DNA. According to the authors' findings, the longer the enzyme goes without locating the GATC sequence of molecules, the less likely that it will undergo this new variation of hopping, but the introduction of a GATC sequence will stimulate the mechanism once again.
According to the paper, hopping can explain the efficiency by which DNA-modifying enzymes can find their recognition sites, despite the presence of an overwhelming amount of non-specific DNA; as well as how enzymes can modify more than one site, despite opposing strand orientations.
The research capitalizes on decades of observation of E. coli's behavior, and factors that contribute to its virulence, or its ability to persist and multiply. Studying the mechanisms that switch these abilities on and off would contribute to how humans can deal with these bacteria, which exist in warm-blooded creatures, but, in certain instances, can cause diseases.
"If we had inhibitors that could prevent the switching, we wouldn't have urinary tract infections, for instance," said Pollak.
The same research group recently reported a similar mechanism in humans, which is disrupted in certain forms of leukemia.
Journal information: Journal of Biological Chemistry
Provided by University of California - Santa Barbara