July 9, 2012 report
Researchers reach new level of clarity in recording proteins moving through lipids

(Phys.org) -- One of the great challenges in biology is recording biological processes at the cellular or even subcellular level. To get a close view, microscopes have to focus on one specific part of whatever it is they are looking at, but because biological processes generally involve movement; that which they are looking at suddenly moves out of the frame. Now it appears a team of international researchers led by Simon Scheuring of Parc Scientifique et Technologique de Luminy, has found a way to record the “dance” that goes on as membrane proteins diffuse within biological membranes. The team describes their technique in their paper published in the journal Nature Nanotechnology.
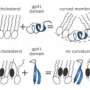
In order for biology to work at the subcellular level, proteins, lipids and sugars along with other molecules have to follow certain rules, which some have likened to dance steps, because some of the most basic rules involve the paring up of molecules. This paring up allows the molecules to carry out activities that cannot be done alone. Also, one of the things that need to happen to sustain life is for proteins to somehow make their way through a layer of lipids, as is the case with Escherichia coli which have an outer membrane coated with proteins that need to make their way inside.
To actually watch the dance that goes on as the proteins try to make their way through the membrane, the researchers built a new kind of microscope apparatus. It involves a stylus, a cantilever and a laser. The tip or stylus of the apparatus moves slowly over the surface of the specimen being studied while the laser serves as a guide. The cantilever allows the stylus to bob up and down in concert with the hills and valleys found below. What this does is allow a camera to record the cellular terrain in similar fashion to a helicopter moving up and down as it surveys a mountain range, trying to maintain the same altitude throughout. By doing so, recordings can be made of very small biological processes as they occur.
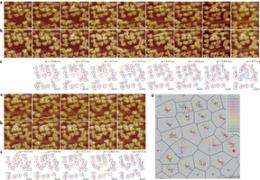
Using this setup, the team was able to record in minute detail the pairing up of molecules and note the way those that weren’t able to pair up made their way through the outer membrane of the E. coli bacteria. The team believes this new technology will open the door to much more research by many teams investigating how biological processes at the subcellular level actually work.
More information: Characterization of the motion of membrane proteins using high-speed atomic force microscopy, Nature Nanotechnology (2012) doi:10.1038/nnano.2012.109
Abstract
For cells to function properly, membrane proteins must be able to diffuse within biological membranes. The functions of these membrane proteins depend on their position and also on protein–protein and protein–lipid interactions. However, so far, it has not been possible to study simultaneously the structure and dynamics of biological membranes. Here, we show that the motion of unlabelled membrane proteins can be characterized using high-speed atomic force microscopy. We find that the molecules of outer membrane protein F (OmpF) are widely distributed in the membrane as a result of diffusion-limited aggregation, and while the overall protein motion scales roughly with the local density of proteins in the membrane, individual protein molecules can also diffuse freely or become trapped by protein–protein interactions. Using these measurements, and the results of molecular dynamics simulations, we determine an interaction potential map and an interaction pathway for a membrane protein, which should provide new insights into the connection between the structures of individual proteins and the structures and dynamics of supramolecular membranes.
Journal information: Nature Nanotechnology
© 2012 Phys.org