Venomous snail key behind therapeutic molecules

Can a painkiller be re-engineered to get a closer look at how proteins bind to communication channels? Researchers across Europe are using state-of-the-art computing techniques to re-engineer a painkiller from the XEP-018 protein, which was identified inside the venom of the Conus consors, a species of sea snail. CONCO, gathering 19 European partners and the J.Craig Venter Institute in the United States, is using the venomous sea snail to develop new therapeutic molecules.
'All senses in our body are transmitted to and from the brain via neurons' quoted Dr Henry Hocking from Utrecht University and a CONCO member. 'The venom of the cone snail has peptides that can disrupt this circuitry. The peptides do this by attaching themselves to the openings in communication channels located on the muscular tissues receiving the neuronal signal. This is sort of like a plug. Once attached, no signal can be transmitted to the brain and you stop feeling pain.'
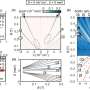
Dr Hocking and colleagues used nuclear magnetic resonance (NMR) to recreate the three-dimensional (3D) structure of the XEP-018 protein, a promising molecule discovered by CONCO members recently described by Philippe Favreau, a researcher at Atheris Laboratories in Switzerland, and colleagues in the British Journal of Pharmacology.
Commenting on the use of NMR, Utrecht's Professor Alexandre Bonvin said: 'NMR is a technique that people might know from hospitals, where magnetic resonance imaging scanners are used. People are put into large magnetic fields and pictures are made of them. In NMR, we put protein molecules inside and bombard them with electromagnetic waves. Instead of making pictures, we measure distances between atoms. If you know all the distances between the atoms, you can try to reconstruct a 3D object of the protein.'
It should be noted, however, that NMR does not indicate the 3D structure of these peptides. The use of computations enables NMR data to be converted into a 3D protein structure.
The team made an NMR analysis on the grid by combining gLite middleware, the next generation middleware for grid computing, with the WENMR e-Infrastructure.
'In order to calculate the 3D structures of proteins, we have to repeat the process many times,' Professor Bonvin was quoted as saying. 'We have to make thousands or tens of thousands of calculations. You get your answer within a couple of hours. WENMR, as a whole, had a submission volume of about 1.5 million jobs last year, corresponding to over 850 central processing unit (CPU) years.'
The Utrecht team is currently assessing whether it is feasible to deploy a dedicated desktop grid within the university. This would help the researchers develop their computational resources even further.
The CONCO consortium is also preparing to initiate the XEP-018 research at the clinical trial stage. 'Many analogues have been designed, synthetized and tested. The product is currently in preclinical development for the treatment of dystonia,' Reto Stöcklin, CONCO scientific team leader and head of the Swiss SME Atheris Laboratories, was quoted as saying. 'Our ultimate goal is to avoid injections and to develop a drug that everybody can use. Using specific devices, such as patches or cell-penetrating peptides to facilitate the penetration of the peptide through the skin, we believe that XEP-018 has great chances of success.'
The researchers pointed out how the WENMR study is giving developers in the global software community the means to share know-how and foster stronger cooperation.
More information:
CONCO: www.conco.eu/index.html
WENMR: www.wenmr.eu/
Utrecht University: www.uu.nl/EN/Pages/default.aspx
Provided by CORDIS