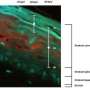
Proteomics identifies targets of ionizing radiation in a human skin model

(Phys.org) -- How better to find out what effect ionizing radiation has on human skin than by using the real thing? Researchers at Pacific Northwest National Laboratory did that by performing a quantitative mass spectrometry study on a reconstituted human skin tissue model to identify areas affected by ionizing radiation exposure. Their results suggest that even very low doses of ionizing radiation activate cell-signaling pathways, resulting in altered protein phosphorylation and possibly altered protein function.
They also demonstrated that the reconstituted human skin system undergoes a coordinated response to maintain tissue integrity and mitigate radiation exposure effects. The results appear in the May 2012 issue of Experimental Dermatology.
"We identified 1052 unique phosphopeptides, 176 of which showed significant changes in response to radiation dose," said PNNL biophysicist Dr. David Stenoien. "This gives us a useful database of phosphorylation sites in the human skin model we used, and the data can be used to assess proteins and molecular pathways affected by radiation."
The researchers know the data doesn't necessarily represent all phosphorylation sites in the system. However, they identified several radiation-sensitive proteins with known and previously unreported roles in radiation responses.
Prominent among these is profilaggrin, a protein important for maintaining skin barrier function and hydration, which undergoes increased phosphorylation at multiple sites. The phosphorylation makes it highly insoluble, and a general decrease in phosphorylation is required before it is digested by proteases. However, to the PNNL team's knowledge, their study provides the first description of these phosphorylation sites in the human linker regions that have the potential to impact profilaggrin processing.
While the effects of high doses of radiation on biological systems and human health are well documented, the health consequences of low dose exposures (≤10 centiGrays or cGy), such as that from a typical full body CT scan, are still being debated. In many cases, the potential health effects of low dose exposures are extrapolated from high dose data because it's been difficult to acquire statistically significant cancer risk data at low doses; therefore, risks may be overestimated. Because humans are exposed continuously to low doses of ionizing radiation from the environment, medical procedures, and in certain occupations, it is important that any health consequences are thoroughly understood.
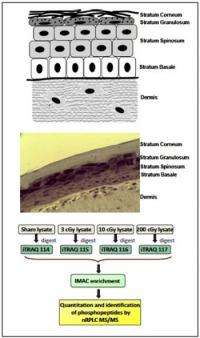
The PNNL team used reconstituted skin tissues, which are metabolically active and recapitulate many skin functions, to study radiation responses in a 3D tissue setting. They applied an isobaric labeling strategy using a sham dose as a control and three doses of radiation (3, 10, and 200 cGy) to identify the 1052 unique phosphopeptides.
This strategy allows multiple samples to be combined within the same multiplexed analysis to quantify the effects of treatment conditions on biological systems. In addition, because the labeling is performed at the post-tryptic peptide level, this method can be used to analyze proteins present in tissues.
This systems biology approach identified multiple novel targets of low dose radiation-activated signaling pathways and demonstrated that the reconstituted human skin system undergoes a coordinated response to maintain tissue integrity and mitigate radiation exposure effects.
Highlighting the need to transition studies into 3D tissues to more accurately recapitulate in vivo responses to low radiation doses, many of the molecular responses involved tissue-level functions. For example, proteins involved in maintaining skin barrier functions and tissue-level defenses to inflammatory agents were found to be important targets of low dose signaling pathways present in differentiated skin tissue.
More information: Yang F, et al. 2012. "Quantitative Phosphoproteomics Identifies Filaggrin and Other Targets of Ionizing Radiation in a Human Skin Model." Experimental Dermatology 23(5):352-357. DOI: 10.1111/j.1600-0625.2012.01470.x
Provided by Pacific Northwest National Laboratory