Taking stock of subsurface microbial communities at Hanford

Taking a census provides valuable information about residents' ages, employment, makeup, living conditions, etc. Most censuses are taken door to door or by mail. But if the community lives in areas that are inaccessible by typical methods, how do you get a meaningful census? That's the question scientists at Pacific Northwest National Laboratory faced when they wanted to tally the microbes within a cross section of the Hanford Site subsurface in south-central Washington state.
It turns out that collaboration is the key. Using gene sequencing provided by the Department of Energy's Joint Genome Institute and samples from a DOE Integrated Field Research Challenge (IFRC) site at Hanford, the scientists were able to obtain the largest census of subsurface microbes taken to date.
The results, which appear in Environmental Microbiology, indicated a region just below the contact between the two major geologic areas of the site that is a zone of active biogeochemical redox cycling. These biogeochemical activities are known to also transform metals such as chromium, uranium, and technetium, which are important contaminants at the site.
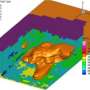
The 1518-km2 Hanford Site is a laboratory unto itself, as its sediments, rock layers, and groundwater host communities of microbes that react with each other and their surroundings. This gives scientists fertile ground-perhaps a better term is layers-for exploring areas as diverse as carbon sequestration and environmental sustainability.
Despite their ecological importance, determining the number and distribution of subsurface microbes has been more difficult than that of other microbial ecosystems. This is because it's hard to retrieve authentic, uncompromised samples from the subsurface.
The site also contains multiple subsurface contaminant plumes that can include metals, radionuclides, nitrate, organic solvents and/or complexing agents. Fundamental science questions about how to best remediate these plumes remain unresolved, which has driven DOE-funded research at the site.
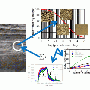
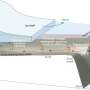
The PNNL team investigated microbial diversity in subsurface sediments at Hanford by analyzing 21 samples recovered from depths of 9-52 m. The samples were collected over distinct geologic strata at the Hanford IFRC site.
They analyzed approximately 8000 nearly full-length 16S rRNA gene sequences across geologic strata that included Hanford formation, Ringold Formation sediments, and the weathered basalt group, with more than 6500 near full-length 16S rRNA gene sequences from Bacteria and 1500 from Archaea.
The scientists characterized the geochemical properties and structure of the microbial community across these different sedimentary formations. The structure and richness of these assemblages of organisms within a specific habitat varied substantially across the geologic strata.
"Some strata had high relative abundances of poorly described bacterial taxa," said Dr. Allan Konopka, PNNL Laboratory Fellow and senior author of the article. "The high numbers of these novel organisms make them likely to catalyze important biogeochemical reactions."
Given the stratigraphic and redox differences between the Hanford formation and the oxic and reduced Ringold units, it was unsurprising to find dissimilar microbial communities in sediments from these zones. However, the scientists also found significant differences in community composition among samples from the same geological stratum, which suggests that these subsurface sediments also comprise diverse microbial habitats, presumably driven by small-scale physical and chemical heterogeneities.
Scientists are not certain which particular organisms catalyze specific redox processes nor what energy sources are driving these reactions. Possible energy sources are organic matter in groundwater or in sediments, iron sulfide minerals in the sediments, or gasses diffusing up from deeper sediment layers. More work is necessary to distinguish among these possibilities.
More information: Lin X, et al. 2012. "Vertical stratification of subsurface microbial community composition across geological formations at the Hanford Site." Environmental Microbiology 14(2):414-425. DOI: 10.1111/j.1462-2920.2011.02659.x
Provided by Pacific Northwest National Laboratory