Scientists reveal how bacteria build homes inside healthy cells

(PhysOrg.com) -- Bacteria are able to build camouflaged homes for themselves inside healthy cells - and cause disease - by manipulating a natural cellular process.
Purdue University biologists led a team that revealed how a pair of proteins from the bacteria Legionella pneumophila, which causes Legionnaires disease, alters a host protein in order to divert raw materials within the cell for use in building and disguising a large structure that houses the bacteria as it replicates.
Zhao-Qing Luo, the associate professor of biological sciences who headed the study, said the modification of the host protein creates a dam, blocking proteins that would be used as bricks in cellular construction from reaching their destination. The protein "bricks" are then diverted and incorporated into a bacterial structure called a vacuole that houses bacteria as it replicates within the cell. Because the vacuole contains materials natural to the cell, it goes unrecognized as a foreign structure.
"The bacterial proteins use the cellular membrane proteins to build their house, which is sort of like a balloon," Luo said. "It needs to stretch and grow bigger as more bacterial replication occurs. The membrane material helps the vacuole be more rubbery and stretchy, and it also camouflages the structure. The bacteria is stealing material from the cell to build their own house and then disguising it so it blends in with the neighborhood."
The method by which the bacteria achieve this theft is what was most surprising to Luo.
The bacterial proteins, named AnkX and Lem3, modify the host protein through a biochemical process called phosphorylcholination that is used by healthy cells to regulate immune response. Phosphorylcholination is known to happen in many organisms and involves adding a small chemical group, called the phosphorylcholine moiety, to a target molecule, he said.
The team discovered that AnkX adds the phosphorylcholine moiety to a host protein involved in moving proteins from the cell's endoplasmic reticulum to their cellular destinations. The modification effectively shuts down this process and creates a dam that blocks the proteins from reaching their destination.
The bacterial protein Lem3 is positioned outside the vacuole and reverses the modification of the host protein to ensure that the protein "bricks" are free to be used in creation of the bacterial structure.
This study was the first to identify proteins that directly add and remove the phosphorylcholine moiety, Luo said.
"We were surprised to find that the bacterial proteins use the phosphorylcholination process and to discover that this process is reversible," he said. "This is evidence of a new way signals are relayed within cells, and we are eager to investigate it."
The team also found that the phosphorylcholination reaction is carried out at a specific site on the protein called the Fic domain. Previous studies had shown this site induced a different reaction called AMPylation.
It is rare for a domain to catalyze more than one reaction, and it was thought this site's only responsibility was to transfer the chemical group necessary for AMPylation, Luo said.
"Revealing that this domain has dual roles is very important to identify or screen for compounds to inhibit its activity and fight disease," he said. "This domain has a much broader involvement in biochemical reactions than we thought and may be a promising target for effective treatments."
During infection bacteria deliver hundreds of proteins into healthy cells that alter cellular processes to turn the hostile environment into one hospitable to bacterial replication, but the specific roles of only about 20 proteins are known, Luo said.
"In order to pinpoint proteins that would be good targets for new antibiotics, we need to determine their roles and importance to the success of infection," he said. "We need to understand at the biochemical level exactly what these proteins do and how they take over natural cellular processes. Then we can work on finding ways to block these activities, stop the infection and save lives."
More information: A paper, Legionella pneumophila SidD is a deAMPylase that modifies Rab1, detailing the work is published in the current issue of the Proceedings of National Academy of Sciences.
ABSTRACT
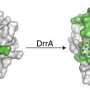
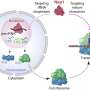
Legionella pneumophila actively modulates host vesicle trafficking pathways to facilitate its intracellular replication with effectors translocated by the Dot/Icm type IV secretion system (T4SS). The SidM/DrrA protein functions by locking the small GTPase Rab1 into an active form by its guanine nucleotide exchange factor (GEF) and AMPylation activity. Here we demonstrate that the L. pneumophila protein SidD preferably deAMPylates Rab1. We found that the deAMPylation activity of Sid D could suppress the toxicity of SidM to yeast and is required to efficiently release Rab1 from bacterial phagosomes. A molecular mechanism for the temporal control of Rab1 activity in different phases of L. pneumophila infection is thus established. These observations indicate that AMPylation-mediated signal transduction is a reversible process regulated by specific enzymes.
Provided by Purdue University