New application makes supercomputing simple
(PhysOrg.com) -- A new open source application developed at Murdoch University is giving researchers a revolutionary new way of accessing supercomputers.
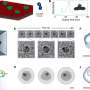
Yabi simplifies supercomputing tasks through a simple web-based workflow environment, essentially replacing the need for complex software programming with a neat, accessible interface.
The web-based application is designed, developed and maintained by the WA Centre of Excellence in Comparative Genomics (CCG), home of the iVEC@Murdoch supercomputer pod.
According to CCG Director Professor Matthew Bellgard, Yabi has the potential to change the way researchers approach scientific endeavours which typically require access to large scale computing and data storage resources.
“Typically, a PhD student in areas such as life science, marine science, atmospheric research and so forth has to learn how to program; they have to know how to install the analysis tools so they can then conduct their detailed data analysis on a supercomputer,” Professor Bellgard said.
“The Yabi system takes away that need for writing scripts and tools and turns the analytic procedures into a simpler drag and drop activity, where scientists can log in, drag tools in and chain them together to create workflows.
“Each of those tools can be running on supercomputers without the need for scientists to have to worry about any of the technical details. In this way scientists can access potentially multiple supercomputing resources in a seamless and transparent fashion via a simple web-based interface.”
Simplifying supercomputing is no small task, and the technologies required to create this kind of interface are fairly young.
“The idea of Yabi has been around for about 12 years,” Professor Bellgard said.
“We’ve been thinking about it for quite some time, but it’s only in the last five years that internet technologies have matured in such a way that we can then leverage them in order to implement a really robust system.
“The system is currently being accessed by scientists around the world and there are now deployments of the Yabi system around the country.”
The CCG works with researchers both to help them improve their Yabi uptake as well as to assist them analyse the massive amounts of results generated.
“They drive their own scientific questions but we can help them with experimental design and data analysis,” Professor Bellgard said.
“We are also working on a number of other open source software projects such as laboratory information management systems and rare disease registries.”
Researchers who use any kind of supercomputing in their work are encouraged to try Yabi – visit ccg.murdoch.edu.au/yabi .
Provided by Murdoch University