TACC supercomputers help researchers find deeper insight into structure and behavior of protein, DNA and RNA
In 1926, Theodor Svedberg won the Nobel Prize in Chemistry for a novel method of separating proteins based on experiments performed on a new device he invented: the analytic ultracentrifuge.
Analytical ultracentrifugation (AUC) experiments spin samples at very high speeds to study how large molecules such as proteins, DNA and RNA, act in solution. Under the influence of centrifugal forces up to 250,000 times as strong as Earth's gravity, materials undergo sedimentation and diffusion processes over time, revealing aspects of the individual molecules' natures.
These processes are essential measurements for biochemists: a way to understand how molecules behave under physiological solution conditions. And 85 years later, scientists are still finding ways to make the analytic ultracentrifuge more useful.
Unlike traditional microscopy where samples are bound to a microscope grid, or x-ray crystallography where they are locked into a crystal with packing forces that may distort the molecule, AUC experiments preserve the native structures and configurations of molecules. They do this by analyzing molecules in solution, where they can dynamically interact and bind to other molecules, or react to environmental changes such as temperature, ionic strength or pH.
.png)
"If you don't have a way to measure your molecule in solution, then a lot of this will escape you," said Borries Demeler, associate professor of biochemistry at The University of Texas Health Sciences Center and director of the Center for Analytical Ultracentrifugation of Macromolecular Assemblies (CAUMA). "By studying biological macromolecules in a solution, it is possible to observe reactions, and follow conformational changes."
AUC is also a very versatile tool to study composition. Even trace amounts of impurities can be resolved by AUC, and mixtures can be analyzed to identify molecular weight and shape distributions.
Initially, the analysis of centrifugal experiments was done manually, but with the emergence of computers and sensors in the 1960s, more precise ways of assessing experimental results were developed. Today's optical systems can follow sedimenting and diffusing molecules by detecting ultraviolet and visible absorption, the refractive index, and fluorescence emission. The signals are captured digitally to allow them to be analyzed by computer.
For more than two decades, Demeler has worked at the intersection of the physical (spinning samples) and the virtual (supercomputer simulations), investigating new methods and developing software to help researchers make the most of their AUC experiments.
As the director of CAUMA, Demeler works with hundreds of investigators around the world, including biophysicists studying the structure and function of biological macromolecules and assemblies, material scientists trying to make more efficient solar cells, and the pharmaceutical industry evaluating the stability of their formulations. As a collaborator in many research projects, he is continually challenged by new research questions and enjoys the interactions with many fascinating scientists.
His largest impact, however, is felt through the creation of the UltraScan software package, and the development of the UltraScan LIMS portal through which researchers analyze their experimental data over the web using advanced computing methods and systems.
"I started writing the very first version of UltraScan using BASIC on a 286 PC back in 1988," Demeler recalled, "and it's gone through many iterations. "
In 2004, Demeler and his colleague, Emre Brookes, began parallelizing the code so it could run on large-scale computer clusters. This dramatically sped up the rate at which samples could be analyzed. It also enabled the researchers to develop high-resolution analysis methods and address an entirely new class of research questions that widened the application of the AUC method.
.png)
UltraScan doesn't just allow researchers to measure the diffusion and sedimentation processes; it decodes the meaning of these processes and uncovers hidden characteristics of the sample.
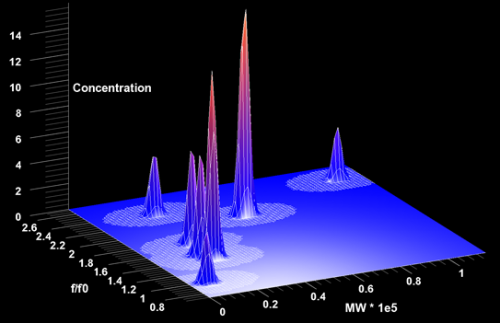
"We often don't know what really is in a solution provided by a collaborator, and we need to get the most out of our analysis," Demeler explained. "To fit the data, we simulate many different components that may be in the solution, and ask the question, ‘How much of each component is present in the actual experiment?'"
This process can be done on a regular computer, but the answers that such a process generates lack the resolution required for clinical or industrial investigations, or simply take too long to complete.
"To squeeze out the last drop of information, you need to go through quite a bit more computational expense," Demeler said. "This is where we kick in with our methods."
UltraScan's numerical methods extract noise, narrow the parameter space, compare multiple experiments, and determine the uncertainty of the result.
While some analyses are performed on a small development cluster in Demeler's lab, the capacity is insufficient to address the most challenging problems, and to satisfy all of the demand for analysis among a growing international group of AUC users. Instead, Demeler relies on the computing systems of the National Science Foundation funded Extreme Science and Engineering Discovery Environment (XSEDE), the most powerful, and robust collection of integrated advanced digital resources and services in the world.
Demeler's simulations use anywhere between 40 and 14,000 processors simultaneously, speeding up the analytic processing by as much as 10,000 times. In 2010-2011, Demeler used 3.5 million computing hours on the Ranger and Lonestar supercomputers at the Texas Advanced Computing Center (TACC) to perform simulations for the open science community.
"It's not just reserved for biochemists and biophysicists," Demeler said. "We might work with a clinician, perform measurements for materials science, or measure the binding strength of a new drug to its target."
Demeler pointed to a recent example of work he is doing with researchers in Germany characterizing fluorescent nanoparticles made out of cadmium telluride crystals for use in solar panels. Using a new detector developed by a collaborator at the Max Planck Institute, he was able to not only measure the hydrodynamic properties, but also observe their individual absorption spectra, and correlate absorbance properties with particle size.
.png)
Whether the application is nanoparticles for industry or biomarkers in blood, AUC together with UltraScan is an incredibly useful tool. But creating the software and algorithms wasn't the final step. Many potential users of AUC and UltraScan are not computer scientists, and Demeler believed a fear of the supercomputer command line interface would prevent them from using the software.
"To get people to adopt this technology, you have to make it easy for them," he said. "It needs to be extremely robust, user-friendly and intuitive."
Through an Advanced Support for TeraGrid Applications (ASTA) grant from the NSF, staff at Indiana University helped Demeler develop a web-based gateway where researchers log in, access their data, and submit jobs as if they were running a very simple web application.
"The user only has to be familiar with the basic analysis procedure and a web browser; familiarity with Unix supercomputing is not required," Demeler said. "Our users really like this approach."
Eighty-five years after its inception, the evolution of the analytic ultracentrifuge continues. The latest challenge involves finding ways to integrate AUC results with results from other solution methods.
Demeler and Brookes are developing an integrated system combining molecular dynamics with hydrodynamic and small angle scattering simulations to screen a large variety of structural conformations against experimental data. This will give a large pool of researchers new insight into the structure and function of molecules under study.
"The knowledge obtained should enhance our understanding of biomolecular processes, including disease processes, which can lead to improved prevention and treatment," said Emre Brookes. "None of this would be feasible without the vast computational resources available through XSEDE."
Demeler and Brookes' long-term dream is to create a way to integrate all known observational methods — including x-ray crystallography, nuclear magnetic resonance imaging, and calorimetry — to see more deeply than we currently do, without losing sight of the natural conditions under which molecules exist.
"It's like taking a picture of an object from many different angles, and every time you take a picture you see something else that adds to the whole," Demeler said. "By combining them all, the new picture will tell you something you didn't know before."
More information: To read more about the challenges of biochemistry and the role of AUC in protein study, see Demeler's recent commentary in the June 2011 edition of Nature Chemical Biology.
The story is courtesy of Faith Singer-Villalobos @ Texas Advanced Computing Center (TACC).
Provided by University of Texas at Austin