Proteins could offer novel antibiotic target
Bacteria are single-celled organisms that inhabit almost every environment on the planet, including the bodies of humans and animals. The cell wall maintains the structural integrity of the cell, and enables the bacteria to survive in its chosen environment. In disease-causing bacteria (pathogens) it also plays a role in the progression of the disease. A group of scientists from Newcastle University and the Nara Institute of Science and Technology in Japan have used Diamond to identify a group of proteins that enable certain bacteria to build effective cell walls. These proteins may provide a novel antibiotic target for a range of disease-causing bacteria.
Gram-positive bacteria include many important human pathogens. In these bacteria, the cell walls contain two major components: peptidoglycan, which is targeted by some of the most successful anti-bacterial compounds and anionic cell wall polymers, which attach to the peptidoglycan. The polymers include wall teichoic acids (WTAs), which play a wide range of roles within the cell including control of autolytic activity, antigenicity and innate immune recognition and antibiotic resistance. How the WTAs physically connect to the peptidoglycan is vital to the final cell wall architecture and how it functions, but currently the enzyme that carries out this important step is unknown.
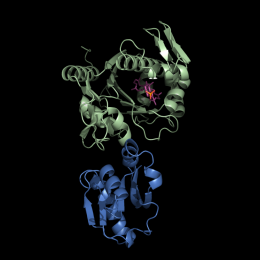
The group have identified a widespread family of proteins, LytR-Cps2A-Psr (LCPs) that could play a role in attaching WTAs to the cell wall. They carried out structural and biochemical analysis of the proteins, including use of Diamond’s MX beamlines. The results suggested that this family of proteins co-ordinate the final stage in the process of connecting the WTA to the bacterial cell wall.
“Studying the final step of bacterial wall synthesis has been very challenging for several reasons. We can’t reconstitute this complex system in vitro, but we have evidence that suggests the LCP family of proteins are the final step in building the cell wall. This discovery, together with structural information on the substrate and some clues to the likely catalytic mechanism, provides us with a highly attractive new target for drug discovery programmes.” said Jon Marles-Wright, Newcastle University.
More information: A widespread family of bacterial cell wall assembly proteins, Yoshikazu Kawai, Jon Marles-Wright, Robert M Cleverley, Robyn Emmins, Shu Ishikawa, Masayoshi Kuwano, Nadja Heinz, Nhat Khai Bui, Christopher N Hoyland, Naotake Ogasawara, Richard J Lewis, Waldemar Vollmer, Richard A Daniel and Jeff Errington, The EMBO Journal, September 2011 doi:10.1038/emboj.2011.358
Provided by Diamond Light Source



















