Biodesign researchers to develop new reagent pipeline for molecular medicine
An ongoing Arizona State University effort to develop a revolutionary class of reagents that holds great promise for the future of medicine has received a major boost with a three-year, $4 million award from the National Institutes of Health.
Scientists John Chaput and Joshua LaBaer in the Biodesign Institute at Arizona State University will lead a new initiative to develop a state-of-the-art pipeline for generating synthetic affinity reagents called "DNA synbodies." The vision is to develop an automated pipeline that will produce low-cost affinity reagents for the entire human proteome, which is the complete ensemble of proteins found in humans.
"Creating a new class of reagents to recognize all of the proteins that are made in our bodies is the next grand challenge in molecular medicine," said ASU professor John Chaput. "The majority of the proteins encoded in our genes have functions that remain unknown. We are making synthetic, affinity reagents to allow for the global study of protein function and how this contributes to disease."
"Proteins provide the verbs to biology; they build, modify, degrade, fold, transport, amplify, proofread, regulate, and the list goes on," said Joshua LaBaer, director of the institute's Virginia G. Piper Center for Personalized Diagnostics. "Most disease is the result of protein dysfunction and nearly all therapeutics target proteins. Nearly all of our methods for studying protein function, as well as for identifying proteins as diagnostics, rely upon the use of specific affinity reagents."
LaBaer points out that the future of medicine will depend on our ability to monitor an individual's health status using personalized diagnostics that will accurately measure proteins as indicators of disease, called biomarkers.
With seed funding from Science Foundation Arizona and the National Cancer Institute, Chaput has developed, characterized and validated the DNA synbody concept. Now, by combining forces with LaBaer, a leader in automated, high-throughput technologies for protein analysis, they will take the DNA synbody concept to the next level and develop the ability to identify affinity reagents that will compete head-to-head with traditional antibodies.
Since the 1970s, the current gold standard is monoclonal antibody technology. But they are made from genetically pure cells and produced in animals via immunization, which is an expensive, time-consuming approach that can take many months of labor. And a single antibody can costs thousands of dollars to produce—with no guarantee the final product, or even commercially available antibodies, will be of high enough quality to recognize their target the same way every time. Even with these limitations, antibodies represent a $5 billion global molecular diagnostics market that focuses on key areas such as infectious disease testing, cancer screening and drug development.
To develop a next generation technology for affinity reagents, LaBaer and Chaput wanted to overcome the shortcomings of current technologies, focusing their efforts on easy-to-assemble building blocks that are cheap, high quality, simple to make, and can be mass produced and easily distributed to labs throughout the world.
"What the scientific community really wants is a technology that's doing it all: faster, cheaper, better, and a technology where you can mass-produce reagents," said Chaput. "Our idea is to create an affinity reagent pipeline that's completely automated using robotics to discover peptides that bind proteins."
Their innovative approach will allow them to identify a select number of affinity reagents from among trillions of possibilities in just a single step. The initial protein targets of the grant are human kinases, important signaling molecules found to participate in many vital metabolic functions of the cell, as well as diseases such as cancer, and are the foundation for new anti-cancer therapeutics. LaBaer's lab has one of the largest collections of human kinases, curating more than 500 genes, and has used this library, called DNASU, to find novel pathways for cancer drug resistance.
"Kinases play such a central role in the how body regulates itself and they affect so many different human diseases that they were a logical place to start," said LaBaer. "But obviously, what excites us the most is that we can apply this approach to any protein that is important to human health. Our lab has a gene collection capable of producing 13,000 of the 20,000 proteins found in humans, so we have a lot of opportunity."
Their ambitious goal by the end of the funding period is to manufacture more than 4,000 new affinity reagents candidates to target against these 100 human kinases, opening up entirely new vistas for cancer research.
"Once this platform is developed, we hope to have demonstrated and validated the versatility of these new reagents and make them available to the greater scientific community and industrial collaborators to advance biomedical research," said Chaput. "We will have the ability to generate affinity reagents for any protein of interest."
The Missing LINC
At the heart of DNA synbody technology lays the ability to imagine entirely new uses for the twisting ladder of the DNA helix.
Chaput refers to DNA synbody assembly technology by a short acronym: ligand interactions by nucleotide conjugates (LINC). This patented technology uses DNA as a programmable scaffold dotted with pairs of peptides around the helix.
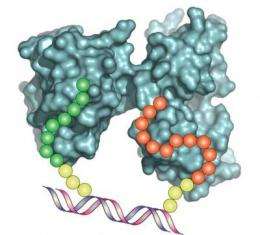
"We construct these molecules in a single-pot reaction," said Chaput. "Our DNA synbodies are unique reagents that use a short DNA backbone to support a pair of specific peptides at defined spacing and angular rotation."
Starting with a vast, peptide library of more than 1 trillion candidates for each protein target, Chaput quickly identifies high affinity peptides in a single high throughput screen. "For each target, the idea is to get 10 good peptides, of which we pick 4 to explore further."
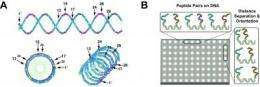
Next, Chaput uses the LINC method to test pairwise combinations of all four peptides on the DNA helix. "In the past, this work was done by a single researcher working on one target at time. Moving forward, we would like to automate this process using state-of-the-art robotics technologies available in the LaBaer lab."
Using liquid handler machines, one peptide is linked to the end of one of the DNA strands, while a second peptide is placed at 10 different locations on a set of complementary strands. The strands are woven together to make a DNA double helix that displays two peptides at a defined distance and angular geometry. The researchers then search the different combinations for synbodies that bind their target protein with high affinity.
"Ideally, we would like to identify multiple DNA synbodies for each target. A major advantage of LINC is the ability to scale the technology against many protein targets," said Chaput. "To our knowledge, no other affinity reagent technology is capable of producing thousands of candidate synbodies."
In addition, Chaput has already made DNA synbodies that recognize and bind proteins better than commercially available antibodies. If the research team is successful, they will not only provide new tools for proteomic research, but also eliminate the 'buyer beware' quality control problems that plague the current gold standard of affinity reagents, monoclonal antibodies.
Provided by Arizona State University