New methods allow for insights into molecular mechanisms of regeneration
Researchers of the Berlin Institute for Medical Systems Biology (BIMSB) at the Max Delbrück Center for Molecular Medicine (MDC) Berlin-Buch have gained new insights into planarian flatworms, which are an attractive model for stem cell biology and regeneration. Close collaboration between four laboratories at the BIMSB led by Stefan Kempa, Christoph Dieterich, Nikolaus Rajewsky and Wei Chen has led to the identification of thousands of gene products, many of which are expressed and are important in stem cell function. This was achieved by precise characterization of all RNA-molecules expressed in the animals' cells, the so-called transcriptome, without using the genome sequence.
Planarians are famous for their almost unlimited ability to regenerate any tissue via pluripotent adult stem cells. Their spectacular regenerative capabilities have been studied for more than 100 years. With the development of new molecular and genetics approaches, planarians have recently re-emerged as a model system for the study of regeneration and stem cells.
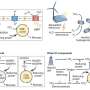
The scientists at the BIMSB combined two existing and complementary sequencing methods to decipher the transcriptome of the planarian Schmidtea mediterranea without depending on genome sequences. Their approach is of great practical importance since the genomes of many organisms are known to be extremely difficult to assemble, even with the current sequencing technologies.
Furthermore, they also were able to identify several novel gene products (mRNAs) of which they proved that they are specifically expressed in the stem cells. It is the first proteomics study of such scale in this phylum (Platyhelminthes), as Wei Chen pointed out. The catalogue of transcripts assembled in their study, together with the identified peptides, dramatically expands and refines planarian research.
More information: De novo assembly and validation of Planaria transcriptome by massive parallel sequencing and shotgun proteomics, Genome Research, July 2011 21: 1193-1200. doi:10.1101/gr.113779.110
Provided by Helmholtz Association of German Research Centres