Researchers decipher protein structure of key molecule in DNA transcription system
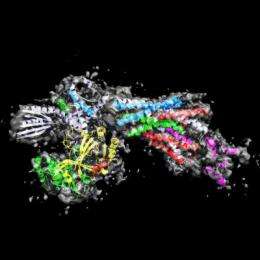
Scientists have deciphered the structure of an essential part of Mediator, a complex molecular machine that plays a vital role in regulating the transcription of DNA.
The research adds an important link to discoveries that have enabled scientists to gain a deeper understanding of how cells translate genetic information into the proteins and processes of life. The findings, published in the July 3 advance online issue of the journal Nature, were reported by a research team led by Yuichiro Takagi, Ph.D., assistant professor of biochemistry and molecular biology at Indiana University School of Medicine.
The fundamental operations of all cells are controlled by the genetic information – the genes –stored in each cell's DNA, a long double-stranded chain. Information copied from sections of the DNA – through a process called transcription – leads to synthesis of messenger RNA, eventually enabling the production of proteins necessary for cellular function. Transcription is undertaken by the enzyme called RNA polymerase II.
As cellular operations proceed, signals are sent to the DNA asking that some genes be activated and others be shut down. The Mediator transcription regulator accepts and interprets those instructions, telling RNA polymerase II where and when to begin the transcription process.
Mediator is a gigantic molecular machine composed of 25 proteins organized into three modules known as the head, the middle, and the tail. Using X-ray crystallography, the Takagi team was able to describe in detail the structure of the Mediator Head module, the most important for interactions with RNA polymerase II.
"It's turned out to be extremely novel, revealing how a molecular machine is built from multiple proteins," said Takagi.
"As a molecular machine, the Mediator head module needs to have elements of both stability and flexibility in order to accommodate numerous interactions. A portion of the head we named the neck domain provides the stability by arranging the five proteins in a polymer-like structure," he said.
"We call it the alpha helical bundle," said Dr. Takagi. "People have seen structures of alpha helical bundles before but not coming from five different proteins."
"This is a completely noble structure," he said.
One immediate benefit of the research will be to provide detailed mapping of previously known mutations that affect the regulation of the transcription process, he said.
The ability to solve such complex structures will be important because multi-protein complexes such as Mediator will most likely become a new generation of drug targets for treatment of disease, he said.
Previously, the structure of RNA polymerase II was determined by Roger Kornberg of Stanford University, with whom Dr. Takagi worked prior to coming to IU School of Medicine. Kornberg received the Nobel Prize in 2006 for his discoveries. The researchers who described the structure of the ribosome, the protein production machine, were awarded the Nobel Prize in 2009. The structure of the entire Mediator has yet to be described, and thus remains the one of grand challenges in structure biology. Dr. Takagi's work on the Mediator head module structure represents a major step towards a structure determination of the entire Mediator.
Provided by Indiana University School of Medicine















