Droplets for detecting tumoral DNA

It will perhaps be possible, in the near future, to detect cancer by a simple blood or urine test. In fact, biologists from CNRS, Inserm, Paris Descartes and Strasbourg universities have developed a technique capable of detecting minute traces of tumoral DNA present in the biological fluids of patients suffering from cancer. The method consists in carrying out ultra-sensitive molecular analyses in microscopic droplets. Successfully tested on genes involved in various cancers, including cancer of the colon and leukemia, it has the potential of becoming a powerful tool for oncologists, both in making a diagnosis and in prescribing a treatment. A clinical study is already envisaged to evaluate this technique. The work is published on the website of the journal Lab on a chip.
When tumoral cells die, they spill their contents into the extracellular medium. These contents, in particular the DNA of cells, are then found in the biological fluids of the patient: blood, lymph, urine, etc. Since the development of most cancers involves genetic factors, a simple blood or urine test could in theory reveal the presence of tumoral DNA and thus cancer as soon as the first cancerous cells die, in other words at a very early stage.
Despite this great promise, there is a snag which explains why physicians cannot yet track down cancers in biological fluids: tumoral DNA is only present in trace amounts in these fluids. In blood, for example, it represents less than 0.01% of the total DNA found in diluted form. However, conventional DNA analysis methods are not sensitive enough to detect such small amounts. Hence the interest of the technique developed by researchers from CNRS, Inserm, the Université de Strasbourg and the Université Paris Descartes, in collaboration with a German team from the Max Planck Institute (Göttingen) and an American company (Raindance Technologies). The considerable advantage of this technique is that it makes it possible to detect DNA thresholds 20,000 times lower than was previously the case in clinics.
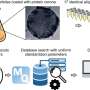
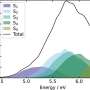
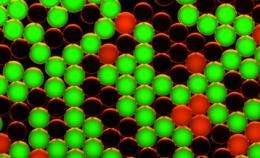
How does it work? A first step consists in distributing the DNA extracted from a biological sample into millions of droplets, which are sufficiently small to contain only a single target gene each. Then, this DNA is amplified by means of modern molecular multiplication methods. Simultaneously, fluorescent molecules specific to each gene interact with the DNA. This key phase provides a sort of gene color code. The droplets are then guided, one by one, into microscopic grooves where they are analyzed by laser: the color of the fluorescent molecules then indicates which gene is present in the droplet. If the droplet emits red fluorescence, for example, the DNA is healthy. If it is green, it is tumoral. If the droplet does not emit any fluorescence, it does not contain the targeted gene. A simple count of the colored spots then makes it possible to determine the tumoral DNA concentration.
The researchers have successfully applied their method to an oncogene (a gene that has the potential of causing cancer) known as KRAS (associated with leukemia and various cancers, such as cancer of the colon, pancreas and lung). The DNA bearing this gene was derived from laboratory cell lines. This new analytical method now needs to be tested in a therapeutic context. A clinical study is already scheduled. If it is a success, physicians will have an efficient “anticancer weapon”, not just for detecting the presence of tumors but also for proposing treatments. The aggressiveness of the cancer, its responsiveness to existing treatments and its risk of recurrence following local treatment: all this information is partly contained in the tumoral DNA. By deciphering it with the microdroplet technology, oncologists could benefit from a powerful diagnostic tool to help predict the evolution of the disease and determine a therapeutic strategy.
More information: Quantitative and sensitive detection of rare mutations using droplet-based microfluidics. Pekin, D., et al., Lab on a chip, published on-line on 19 May 2011, DOI:10.1039/C1LC20128J
Provided by CNRS