New technique tracks viral infections, aids development of antiviral drugs
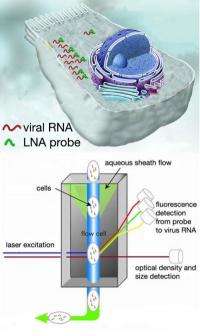
Scientists at the Naval Research Laboratory Center for Bio-Molecular Science and Engineering have developed a method to detect the presence of viruses in cells and to study their growth. Targeting a virus that has ribonucleic acid (RNA) as its genetic makeup, the new technique referred to as locked nucleic acid (LNA) flow cytometry-fluorescence in situ hybridization (flow-FISH), involves the binding of an LNA probe to viral RNA.
While individual parts of the technique have been developed previously, Drs. Kelly Robertson and Eddie Chang, in collaboration with researchers at the NRL Lab for Biosensors and Biomaterials, demonstrate for the first time that the combination of LNA probes with flow-FISH can be used to quantify viral RNA in infected cells. This also allows the scientists to monitor the changes in viral RNA accompanying antiviral drug treatment.
Once the probe is bound to the viral RNA inside mammalian cells, it is tagged with a fluorescent dye, then thousands of these tagged cells are measured rapidly by "flow cytometry" — a method for counting and examining microscopic particles, such as cells and chromosomes, by suspending them in a stream of fluid and passing them by an electronic detection apparatus.
"The ability to rapidly measure thousands of cells for the presence of virus, sets this technique apart from currently used methods to monitor viral replication," said Robertson.
Traditionally, antibodies used to detect viruses must be produced and calibrated for each specific strain and are highly susceptible to viral mutations. Assays commonly used for quantifying viral loads and for drug development can be time consuming and rely on visible signs of cell damage, which is not produced in all viruses and can take long periods of time to occur.
Techniques such as quantitative reverse transcription-polymerase chain reaction (qRT-PCR), microarrays, and enzyme-linked immunosorbent assays (ELISAs), while highly sensitive, involve the lysis [the breaking down] of cells prior to measurement and are therefore unable to provide information about cellular viability, infected cell phenotypes, percentage of infected cells or the variation in infection among a cell population. The LNA probe differs from traditional nucleotide probes by binding more tightly to its target RNA.
LNA-flow FISH presents a fast and easy way to screen for compounds with antiviral activity and could be adapted for monitoring infections in the blood for vaccine therapy and development. This method adds a necessary tool for several emerging areas in cell biology that enables the use of high throughput measurements for entire populations and improves statistical analyses.
"This method can be expanded by adding more than one kind of LNA probe to enable multiple detection of different viral and host RNA," adds Robertson. "The multiplexing enhancement can be used to better understand infectious agents, allowing this technique to be used to aid in the development of antiviral drugs for a variety of viruses."
LNA flow-FISH offers an advantage over other techniques due to its simplicity and superiority. Methods involving genetic recombination of the virus to express a fluorescent protein as a means to mark the presence of virus can utilize flow cytometry for large-batch analysis of infected cells. However, an exception to this approach is viral strains that have not acquired genetic mutations, known as wild-type viruses (such as strains of Human Immunodeficiency Virus-HIV), which would require a large initial investment of labor for engineering each virus of interest.
Provided by Naval Research Laboratory