Biosensors: A handy kit
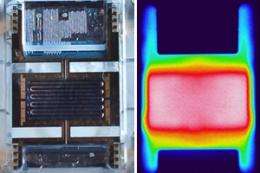
(PhysOrg.com) -- A silicon-based microfluidic chip that distinguishes different viral strains shows potential for the quick on-site diagnosis of infectious diseases.
The control of infectious diseases such as the 2009 H1N1 pandemic influenza hinges on handy analytical tools that can rapidly and accurately identify infected patients at the doctor’s office or at an airport. For this reason, there has been much interest in technologies that could enable replacement of the bulky instruments used at present with point-of-care testing devices. Linus Tzu-Hsiang Kao and co-workers at the A*STAR Institute of Microelectronics and the Genome Institute of Singapore have now developed a silicon-based microfluidic system that is able to sense and differentiate the H1N1 virus from other seasonal influenza strains in ultrasmall specimens.
The detection and characterization of viral strains is now routinely performed using an assay method called real-time reverse transcription polymerase chain reaction (RT-PCR), a method that typically calls for specialized laboratory instruments and skilled personnel. Kao’s team, however, was able to integrate the PCR function into a compact two-module microfluidic chip using standard semiconductor technology. “The system will be suitable for use as a portable diagnostic tool for on-the-spot screening of highly contagious viruses, such as the influenza A H1N1 strain,” says Kao.
Because untreated influenza samples usually contain minute amounts of viral RNA mixed with other nucleic acids and proteins, the researchers designed an ‘on-chip’ PCR module that amplifies target sequences for both H1N1 and seasonal viruses at the same time. The key to their compact screening technology, however, is the silicon-nanowire sensing module used for virus identification. The nanowires in the module are modified with nucleic acid-containing polymers that specifically bind the target DNA, which results in a change in electrical resistance in proportion to the concentration of target DNA present in the sample.
The team fabricated the PCR module, which includes a reaction chamber connected to small aluminum heaters and temperature sensors through tiny channels, directly into a silicon chip using an etching technique. They then constructed the silicon nanowires by optical lithography and finally immobilized the nucleic acid-containing polymers.
Experiments revealed that the small size of the PCR chamber gave it a uniform temperature distribution (see image), providing an ideal environment for efficient RNA amplification. The PCR module also responded much faster to heating/cooling cycles than standard instruments because of the small sample volume—leading to quicker diagnoses.
The team is currently planning to improve the sample extraction module. “We are in the process of building a fully automated and integrated prototype, which will allow us to proceed to clinical validation with our collaborators,” says Kao.
More information: Kao, L. T.-H. et al. Multiplexed detection and differentiation of the DNA strains for influenza A (H1N1 2009) using a silicon-based microfluidic system. Biosensors and Bioelectronics 26, 2006–2011 (2011).
Provided by Agency for Science, Technology and Research (A*STAR)