Bacteria have evolved a unique chemical mechanism to become antibiotic-resistant
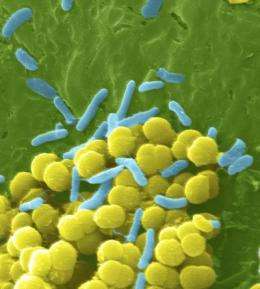
For the first time, scientists have been able to paint a detailed chemical picture of how a particular strain of bacteria has evolved to become resistant to antibiotics. The research is a key step toward designing compounds to prevent infections by recently evolved, drug-resistant "superbugs" that often are found in hospitals, as well as in the general population.
A paper describing the research, by a team led by Squire Booker, an associate professor in the department of chemistry and the department of biochemistry and molecular biology at Penn State University, will be posted by the journal Science on its early-online Science Express site on 28 April. This paper is a continuation of research led by Booker published in another paper in Science earlier this month.
The team began by studying a protein made by a recently evolved "superbug." Booker explained that, several years ago, genetic studies had revealed that Staphylococcus sciuri -- a non-human bacterial pathogen -- had evolved a new gene called cfr. The protein created by this gene had been found to play a key role in one of the bacterium's mechanisms of antibiotic resistance. Later, the same gene was found to have crossed over into a strain of Staphylococcus aureus -- a very common kind of bacteria that constitutes part of the flora living in the human nose and on the skin, and which is now the cause of various antibiotic-resistant infections. Because this gene often is found within a mobile DNA element, it can move easily from a non-human pathogen to other species of bacteria that infect humans. "The gene, which has been found in Staphylococcus aureus isolates in the United States, Mexico, Brazil, Spain, Italy, and Ireland, effectively renders the bacteria resistant to seven classes of antibiotics," Booker explained. "Clearly, bacteria with this gene have a distinct evolutionary advantage. However, until now, the detailed process by which the protein encoded by that gene affected the genetic makeup of the bacteria was unclear; that is, we didn't have a clear 3D picture of what was going on at the molecular level."
To solve the chemical mystery of how such bacteria outsmart so many antibiotics, Booker and his team investigated how the Cfr protein accomplishes a task called methylation -- a process by which enzymes add a small molecular tag to a particular location on a nucleotide -- a molecule that is the structural unit of RNA and DNA. When this molecular tag is added by a protein called RlmN, it facilitates the proper functioning of the bacterial ribosome -- a gigantic macromolecular machine that is responsible for making proteins that bacteria need to survive. Many classes of antibiotics bind to the ribosome, disrupting its function and thereby killing the bacteria. The Cfr protein performs an identical function as the RlmN protein, but it adds the molecular tag at a different location on the same nucleotide. The addition of the tag blocks binding of antibiotics to the ribosome without disrupting its function. "What had perplexed scientists is that the locations to which RlmN and Cfr add molecular tags are chemically different from all others to which tags routinely are appended, and should be resistant to modification by standard chemical methods," Booker said. "What we've discovered here is so exciting because it represents a truly new chemical mechanism for methylation. We now have a very clear chemical picture of a very clever mechanism for antibiotic resistance that some bacteria have evolved."
Booker also said he believes the next step will be to use this new information to design compounds that could work in conjunction with typical antibiotics. "Because we know the specific mechanism by which bacterial cells evade several classes of antibiotics, we can begin to think about how to disrupt the process so that standard antibiotics can do their jobs," he said.
Provided by Pennsylvania State University