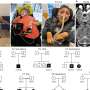
Newly identified RNA sequence is key in microRNA processing
Researchers at Tufts University School of Medicine and Tufts Medical Center have identified an RNA sequence that promotes increased numbers of specific microRNAs (miRNAs), molecules that regulate cell growth, development, and stress response. The discovery helps researchers understand the links between miRNA expression and disease, including heart disease and cancer. The findings are published in the August 13 issue of Molecular Cell.
"A growing body of evidence shows that abnormal expression of miRNAs can contribute to human diseases such as heart disease and cancer. A better understanding of how miRNAs are generated and how they regulate genes may provide important insights into the mechanisms of physiological disorders such as heart disease and cancer," said senior author Akiko Hata, PhD, associate professor in the department of biochemistry at Tufts University School of Medicine (TUSM) and a member of the biochemistry and cell, molecular and developmental biology program faculties at the Sackler School of Graduate Biomedical Sciences at Tufts.
MiRNAs are initially formed as a long sequence of RNA called the primary miRNA. This molecule undergoes several steps to transform it into mature miRNA. Once formed, the mature miRNAs regulate gene expression by silencing or activating target genes. More than 700 human miRNAs with various functions are currently known.
Hata and colleagues previously found that the processing of some miRNAs could be regulated in response to cellular signals from a specific signaling pathway. In the current study, Hata and colleagues found that most of the miRNAs regulated by this signaling pathway share a common RNA sequence. When this RNA sequence was mutated, the signaling pathway no longer regulated miRNA processing. Conversely, when the RNA sequence was introduced into a new miRNA, the miRNA became responsive to the signaling pathway.
"An enzyme called Drosha is needed for miRNA processing. Our previous studies determined that proteins called Smads are also required for the processing of some miRNAs in response to cellular signals. Now, we have identified the RNA sequence that recruits Drosha and Smads for miRNA processing in response to the signaling pathway," said first author Brandi Davis, PhD, a 2010 graduate of the biochemistry program at the Sackler School and a postdoctoral fellow in Hata's lab. "We knew that Smad proteins regulate gene expression by binding to DNA. Our current study is exciting because it shows that Smads play an additional role, controlling miRNA expression by binding to the structurally different RNA."
While miRNAs were first discovered in 1993, scientists did not link them to gene regulation until nearly ten years later. Now, scientists are working to understand how miRNA expression is controlled, what genes miRNAs target, and how varying levels of miRNAs are related to human disease, particularly heart disease and cancer.
"Scientists are just beginning to understand the roles of miRNA in the body, and this study adds another piece to the puzzle. By investigating the mechanisms that govern which genes are translated and which genes are silenced, we can begin to understand how miRNAs impact the progression of cardiovascular diseases and cancer," said Hata.
Hata is also the director of the Molecular Signaling Laboratory in the Molecular Cardiology Research Institute (MCRI) at Tufts Medical Center. The MCRI, with investigators and physician-scientists from Tufts University School of Medicine and Tufts Medical Center, is dedicated to the study of the molecular mechanisms of human cardiovascular disease, the translation of bench findings to new bedside strategies for diagnosis and therapy, and the mentoring of MD and PhD trainees committed to a career in academic cardiovascular research.
More information: Davis BN, Hilyard AC, Nguyen PH, Lagna G, Hata A. Molecular Cell. 2010. (August 13); 39: 373. "Smad Proteins Bind a Conserved RNA Sequence to Promote MicroRNA Maturation by Drosha." Published online August 12, 2010, doi: 10.1016/j.molcel.2010.07.011