Nanofluidics Identify Epigenetic Changes One Molecule at a Time
(PhysOrg.com) -- Using a system of nanofluidic channels and multicolor fluorescence microscopy, a team of investigators at Cornell University has developed a method that analyzes the binding of DNA and DNA-binding proteins known as histones at specific locations along individual DNA molecules. The data generated using this method provides information on the so-called epigenetic state of a cell, which reflect differences in the genes that a given cell is expressing at any one time.
This research effort was led by Paul Soloway, Ph.D., and Harold Craighead, Ph.D., who is also the principle investigator of the Cornell University Physical Sciences-Oncology Center, one of eight newly established centers funded by the National Cancer Institute to identify and study the physical and biological laws and principles that guide the development and spread of cancer. The investigators published the results of this project in the journal Analytical Chemistry.
Every cell in the body contains the same genetic blueprint, but what differentiates a liver cell from a heart cell is a series of DNA modifications, such as methylation, that determines the specific set of genes that are expressed in a specific type of cell. These modifications are known as epigenetic, rather than genetic, changes since they don't alter DNA's sequence, just its structural properties. Those structural changes determine which genes are accessible to the many proteins involved in turning genetic information into specific proteins.
There are many techniques that researchers can use to probe such epigenetic changes, but these methods require large numbers of cells, and thus, produce an average picture of epigenetic state. In addition, these techniques cannot survey the entire genome, nor can they examine two different types of epigenetic changes simultaneously.
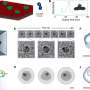
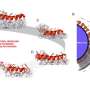
To solve these limitations, the Cornell team created a nanofluidic device capable of flowing individual DNA molecules through a channel and past a detector that can record and analyze the fluorescence of DNA and its associated proteins in real time. The researchers also demonstrated that they can take DNA stripped of its proteins, label it with a fluorescent molecule that binds to methylated bases, and detect specific locations of DNA methylation.
In this set of experiments, the researchers used their nanofluidic system to reveal the frequency and coincidence of epigenetic changes in single DNA molecules. The investigators believe, however, that they will be able to modify the device to rapidly sort DNA-protein structures based on their epigenetic signatures. The sorted chromatin fragments could then be studied further using all the tools of DNA, including DNA sequencing.
his work is detailed in a paper titled, "Single Molecule Epigenetic Analysis in a Nanofluidic Channel." An abstract of this paper is available at the journal's Web site.
Provided by National Cancer Institute