How Botulism Paralyzes Nerve Cells: New Details Revealed
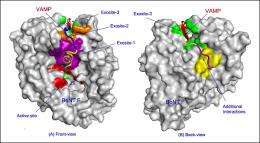
(PhysOrg.com) -- New structures of a botulism toxin interacting with a mimic of the nerve-cell protein it destroys suggest new ways to block this often-fatal interaction. Indeed, the mimic molecules have such high affinity for the toxin and bind to it so tightly that they themselves could possibly serve as anti-toxin drugs with further modification, the researchers said.
The atomic-resolution structures were made at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory in collaboration with scientists from the U.S. Army Medical Research Institute for Infectious Diseases, and will be published online on June 21, 2009, in Nature Structural & Molecular Biology.
Botulism toxins are among the deadliest known poisons. Minute quantities in improperly canned goods can cause a fatal form of food poisoning. Recently, some forms have been used in medical settings to smooth facial wrinkles and to quell bladder spasms to stem urinary leakage. But fear of their use as a bioterror weapon has made the toxins notorious — and the push for developing antitoxin drugs or vaccines a high priority.
The toxins come in seven distinct varieties, but all work the same way: One portion of the toxin binds to a nerve-cell membrane; another portion moves a smaller “catalytic domain” into the cell; then this catalytic domain binds to and cleaves a nerve-cell protein, making it impossible for the nerve cell to “fire,” or send signals. The result is paralysis — and often, death.
“This study looked specifically at how the catalytic domain of one type of neurotoxin, neurotoxin F, recognizes and binds to its target nerve-cell protein to perform this final, paralyzing step,” said Brookhaven Lab biologist Subramanyam Swaminathan, who led the research team.
The team first synthesized two different mimics of the target nerve-cell protein. They then allowed each to bind to the catalytic domain of the toxin, and analyzed the structures using high-intensity x-rays at Brookhaven’s National Synchrotron Light Source (NSLS). Analyzing how the x-rays bounce off the structure allows scientists to reconstruct extremely high-resolution, 3-D images showing the positions and relative orientations of the atoms making up the proteins.
“Our structures reveal that portions of the toxin that are distant from the ‘active site’ that cleaves the nerve-cell protein are crucial to the toxin’s ability to bind to and destroy this protein,” Swaminathan said. Biochemical analysis confirmed the existence and importance of these “exosites,” further validating the crystal structures.
“Because these exosites play such an essential role in the toxin’s ability to bind to and cleave the nerve-cell protein, they could serve as additional targets for the development of drugs designed to interfere with the toxin’s deadly action,” Swaminathan said.
The scientists are also exploring the possibility that the inhibitor molecules they used in this study as mimics for the nerve-cell protein could themselves serve as anti-toxin drugs.
“These inhibitors are attractive candidates for anti-botulinum drug development,” Swaminathan said. “To do so, we’d need to find a way for the inhibitor to reach the toxin inside nerve cells.” One possibility would be to add a transmembrane sequence or some other means of intracellular transport to the inhibitor molecule.
Work on all seven variations of the toxin is essential to understanding common mechanisms that may aid in the design of drugs that work across several different types, or ideally, broadly against all seven. Swaminathan’s group has studied six of the seven varieties. (See related links, below.)
“The mere existence of a vaccine or anti-toxin drugs would help mitigate the extreme fear of a bioterror attack,” Swaminathan said.
Understanding the detailed structures of the toxins and how they interact with their target proteins could also lead to advances in the ways they can be used safely in a medical setting.
Provided by Brookhaven National Laboratory (news : web)