Virtual human puts HIV drug to test
Harnessing the power of supercomputing, ‘grid’ technology and using a so-called ‘virtual physiological human’ (VPH), European researchers have simulated how well an HIV drug blocks a key protein in the lethal virus. The days of trial and error could be numbered.
Thanks to the power of supercomputing, scientists in the UK have shown an early example of the virtual physiological human in action. Carried out earlier this year, the method could pave the way to personalised drug treatment, such as for HIV patients developing resistance to their current regimes.
The human body is too complex to replicate using a single computer or even several computers strapped together. To fully simulate our inner workings, the VPH has to link networks of computers nation- and worldwide. With all this power assembled, scientists can then carry out studies of "supercomputing" proportions, such as the effects of a drug at the organ, tissue, cell and even molecular levels.
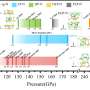
A team from University College London (UCL) in the UK ran simulations to predict how strongly the HIV-inhibiting drug saquinavir would bind to three versions of a viral protein called HIV-1 protease. The protein is used by the virus to propagate itself and, in mutated forms, associated with resistance to the antiretroviral saquinavir. The results are published in the Journal of the American Chemical Society.
Saquinavir is just one of a number of drugs designed to block HIV-1 protease. Currently, doctors have no way to match the drugs to the profile of the virus as it changes in each patient. ‘Trial and error’ is the only solution. With VPH, doctors would be able to see which drugs would be most effective for any given patient.
Borrowed supercomputing
Team leader Professor Peter Coveney of UCL says the study is a first step towards the ultimate goal of “on-demand” medical computing, where doctors could one day “borrow” supercomputing time from national grids to make critical decisions on life-saving treatments.
“For an HIV patient, a doctor could perform an assay to establish the patient’s genotype and then rank the available drugs’ efficacy against that patient’s profile based on a rapid set of large-scale simulations, enabling the doctor to tailor the treatment accordingly,” he offers as an example.
But the professor concedes that the sheer computing power needed to run these simulations is huge. In this latest study, the trials had to be carried out across several supercomputers running off both the UK’s National Grid Service and the US TeraGrid. The work took two weeks and used the same amount of computing power as that needed to perform a long-range weather forecast.
“We have some difficult questions ahead of us, such as how much of our computing resources could be devoted to helping patients and at what price,” says Coveney. “At present, such simulations … might prove costly for the UK National Health Service, but technological advances and those in the economics of computing would bring costs down.”
EU support for the study came from the ViroLab (‘Virtual laboratory for decision support in viral disease treatment’) project. Coveney and his team are now looking at all the protease inhibitor drugs in the same way. A new EU-funded VPH initiative will have €72 million at its disposal to boost collaboration between clinicians and scientists exploring patient-specific medical treatments based on modern modelling and simulation methods.
Source: ICT Results