Massive reanalysis of genome data solves case of the lethal genes
It is better to be looked over than overlooked, Mae West supposedly said. These are words of wisdom for genome data-miners of today. Data that goes unnoticed, despite its widespread availability, can reveal extraordinary insights to the discerning eye. Such is the case of a systematic analysis by the U.S. Department of Energy Joint Genome Institute (DOE JGI) of the massive backlog of microbial genome sequences from the public databases.
The survey identified genes that kill the bacteria employed in the sequencing process and throw a microbial wrench in the works. It also offers a possible strategy for the discovery of new antibiotics. These findings are published in the Oct. 19 edition of the journal Science.
In nature, promiscuous microbes share genetic information so readily that using genes to infer their species position on the evolutionary tree of life was thought to be futile. Now, researchers at DOE JGI have characterized barriers to this gene transfer by identifying genes that kill the recipient bacterium upon transfer, regardless of the type of bacterial donor. These lethal genes also provide better reference points for building phylogenic trees—the means to verify evolutionary relationships between organisms.
“At DOE JGI, we are responsible for producing and making publicly available genomes from hundreds of different microbes, most of which are relevant to advancing the frontiers of bioenergy, carbon cycling, and bioremediation,” said Eddy Rubin, DOE JGI Director. “We realized that sequencing a genome is like conducting a massive experiment in gene transfer. By checking which genes could not be sequenced, we discovered barriers to transfer.”
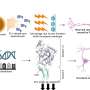
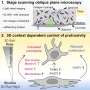
The industrial-scale “shotgun” DNA sequencing strategy typically involves sheering the organism’s DNA into manageable fragments, and then inserting these fragments into a disarmed strain of E. coli, which is used as an enrichment culture—to grow up vast amounts of the target DNA. The team led by Rubin showed that this sequencing process mimics the transmission of DNA from one organism to another, a mechanism called horizontal gene transfer. This phenomenon occurs in nature, allowing one organism to acquire and use genes from other organisms. While this is an extremely rare event in animals, it does occur frequently in microorganisms and is one of the main sources for the rapid spread of antibiotic resistance among bacteria.
“When you sequence a genome, you never get the whole genome reconstructed in one pass,” said Rubin. “You always get gaps in the assembly. This is annoying, expensive, and compels us to close the gaps and finish the puzzle so that we could tell the story behind the sequence. Our breakthrough was in understanding that gaps occur because some genes cannot be transferred to E. coli—because they are lethal.”
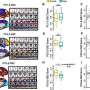
So Rubin and his colleagues sifted through more than nine billion nucleotides to assess gaps in 80 different genomes. They found that the same genes, over and over again, caused these gaps, meaning that they could not be transferred into the E. coli.
“We use the bits that people usually throw away, the gaps of information keeping us from finishing an assembly,” Rubin said. “We identified a set of genes that, if you add another copy or you tweak its expression, the host dies.
“The genes we categorized, while providing us a lesson in the evolutionary history of the organism, now suggest a short-cut for finishing genomes,” Rubin said. “In addition, it offers a new strategy for screening molecules that may represent the next generation of broad-spectrum antibiotics. We expect that many organisms, not just E. coli, are susceptible to being killed if they take up certain genes that are over-expressed. We have strong evidence that most microbes behave like that.”
Source: DOE/Joint Genome Institute