How to steer a moving cell
Researchers at the University of California, San Diego (UCSD) School of Medicine have developed new technology which, combined with proteomics – the large-scale study of the structure and function of proteins and their functions – has allowed them to map an extensive network of the signaling proteins that control cell movement.
Their work, providing the first comprehensive profile of cell movement, could lead to a better understanding of cell migration in cancer metastasis and inflammatory disease. The study will be published the week of May 7-11 in the journal Proceedings of the National Academy of Sciences.
Extra-cellular messengers called chemokines are families of small proteins secreted by cells that regulate the cells’ directional movement, or chemotaxis. Cells possess an innate ability to migrate, an inner compass that somehow senses the presence of chemokines. But in metastasis, the cell’s inner compass goes awry – allowing cells to leave the primary tumor, crawl through tissues and enter blood vessels – spreading the cancer throughout the body.
Richard Klemke, Ph.D., professor of pathology at UCSD School of Medicine and the Moores Cancer Center, and his colleagues set out to better understand the complexity of signaling mechanisms within the cell that become de-regulated and allow cells which are usually static to begin migrating.
The researchers hope to fully define the protein components of the compass to gain a better understanding of what directs cell migration or, in the case of cancer cells and inflammation, cause cells to migrate where they normally wouldn’t.
"The ability to spatially organize specific groups of signaling proteins to the front or back of the cell is what drives cell polarization and directional movement," said Klemke. "It is the steering wheel of the cell." Now the researchers want to determine how the large numbers of signaling molecules that make up the compass are functionally integrated to steer the cell under normal and pathological conditions.
"A surprising finding was that many of the proteins identified in the neuronal network, the ‘wiring diagram’ that controls early development of neural networks in the embryo, are also found to control the movement of normal and cancerous cells," said Klemke. "This is apparently a fundamental, evolutionarily conserved process in migrating cells. So it clearly has an important purpose to the cell."
A network of proteins called the cytoskeleton, the cell’s internal scaffolding, determines cell shape, helping the cell to grow and develop "growth cones." These are specialized structures at the tips of growing nerve fibers, called axons, which sense guidance signals in their environment and "steer" the axons. Cell movement also requires polarization to position the cell and help navigate its movement. Polarization is characterized by formation of a leading "false foot" or pseudopodium and a trailing rear "foot" at the back called a cell body, which is detached in the process of locomotion.
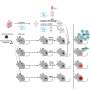
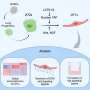
In order to understand the inner workings of the cell’s "steering wheel," the researchers developed a method to cross section the front and back of the chemotactic cell in order to analyze the components of its protein network. The cell fractionation equipment developed in Klemke’s lab is the first method to enable researchers to study the entire network of signaling proteins.
"To find out what goes wrong during metastasis, we need to understand how the signaling networks are controlled in normal healthy cells," said Klemke. "This is the first time a group of researchers from several disciplines – biologists, chemists, proteomics researchers and computational biologists who can integrate large data sets – have applied a global approach to analyzing how proteins regulate cell movement."
The research team’s profile of chemotactic cells is the first and most comprehensive catalog of proteins that exists to date, according to Yingchun Wang, a post-graduate researcher in the Klemke lab.
Source: University of California - San Diego